Page 4 - Voda_Dapporto_alii_2016
P. 4
www.nature.com/scientificreports/
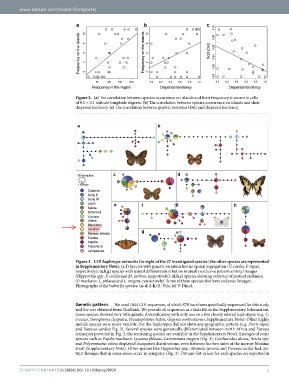
Figure 2. (a) The correlation between species occurrence on islands and their frequency at source in cells
of 0.1 × 0.1 latitude-longitude degrees. (b) The correlation between species occurrence on islands and their
dispersal tendency. (c) The correlation between genetic variation (Dst) and dispersal tendency.
Figure 3. COI haplotype networks for eight of the 27 investigated species (the other species are represented
in Supplementary Note). (e,f) Species with genetic variation but no spatial segregation (V. cardui, P. rapae,
respectively); (a,b,g) species with spatial differentiation but no mutually exclusive pattern among lineages
(Hipparchia spp., P. cecilia and M. jurtina, respectively); (d,h,c) species showing evidence of mutual exclusion
(P. machaon, L. phlaeas and L. megera, respectively). Some of these species also have endemic lineages.
Photographs of the butterfly species: (a–d, f–h) R. Vila; (e) V. Dincă.
Genetic pattern. We used 1044 COI sequences, of which 878 have been specifically sequenced for this study
and the rest obtained from GenBank. We provide all sequences as a fasta file in the Supplementary Information.
Some species showed very little genetic diversification with only one or a few closely related haplotypes (e.g. C.
croceus, Gonepteryx cleopatra, Pseudophilotes baton, Gegenes nostrodamus; Supplementary Note). Other highly
mobile species were more variable, but the haplotypes did not show any geographic pattern (e.g. Pieris rapae
and Vanessa cardui; Fig. 3). Several species were genetically differentiated between north Africa and Europe
(examples provided in Fig. 3, the remaining species are available in the Supplementary Note). Lineages of some
species such as Papilio machaon, Lycaena phlaeas, Lasiommata megera (Fig. 3), Carcharodus alceae, Aricia spp.
and Polyommatus celina displayed chequered distributions, even between the two sides of the narrow Messina
strait (Supplementary Note). Other species like Hipparchia spp., Maniola jurtina and Pyronia cecilia had dis-
tinct lineages that in some areas occur in sympatry (Fig. 3). Dst and Gst values for each species are reported in
Scientific RepoRts | 6:28828 | DOI: 10.1038/srep28828 4